


Hello! 👋 I am a scientist-founder building AI systems for chemistry and scientific discovery. I am a co-founder, CTO, and chief scientist at molecule.one. My background is in deep learning, and I have co-authored widely cited research on the foundations of neural networks. I also serve the research community as an action editor at TMLR and as an area chair for leading conferences, most recently ICLR 2026.
At molecule.one, I led the development of MARIA, a first-of-its-kind microliter high-throughput chemistry platform. We named it after Maria Skłodowska-Curie to reflect our ambition: to automate chemistry and expand the pace and scope of scientific discovery. We built the largest microliter reaction dataset and used it to train models that, for the first time, surpassed human-level accuracy, supporting multiple commercial projects today.
I am interested in learning and optimization across natural and artificial systems, from deep neural networks to the scientific process and the economy. Today, I apply this perspective to autonomous discovery in chemistry at Molecule.one.
I completed a postdoc at New York University with Kyunghyun Cho and Krzysztof Geras, and before that a PhD at Jagiellonian University focused on the foundations of deep neural networks. Along the way, I collaborated with researchers at Mila (Yoshua Bengio), the University of Edinburgh, and Google Research.
During my PhD, I studied the foundations of gradient-based training and co-authored work showing how optimization dynamics can drive neural networks into high-curvature regimes, helping explain the role of learning rate in generalization and training stability, including an ICLR 2020 spotlight paper and subsequent discussion in Understanding Deep Learning (MIT Press).








For a full list, please see my Google Scholar profile.
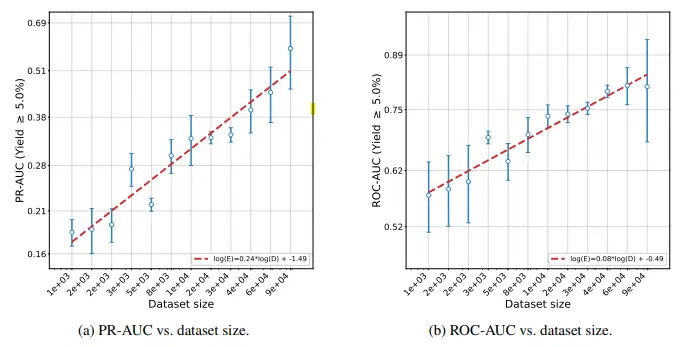
M. Sadowski, L. Sztukiewicz, M. Wyrzykowska, [...], S. Jastrzebski
NeurIPS 2025 Workshop AI4Science
paper
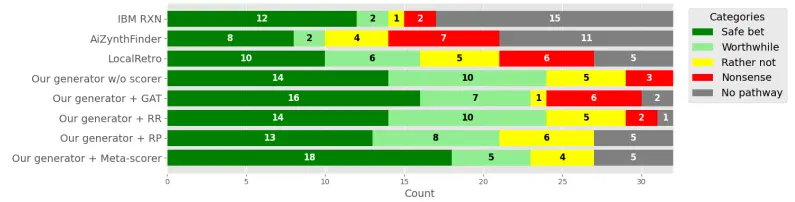
M. Sadowski, [...], S. Jastrzebski
NeurIPS 2025 Workshop AI4Science
paper
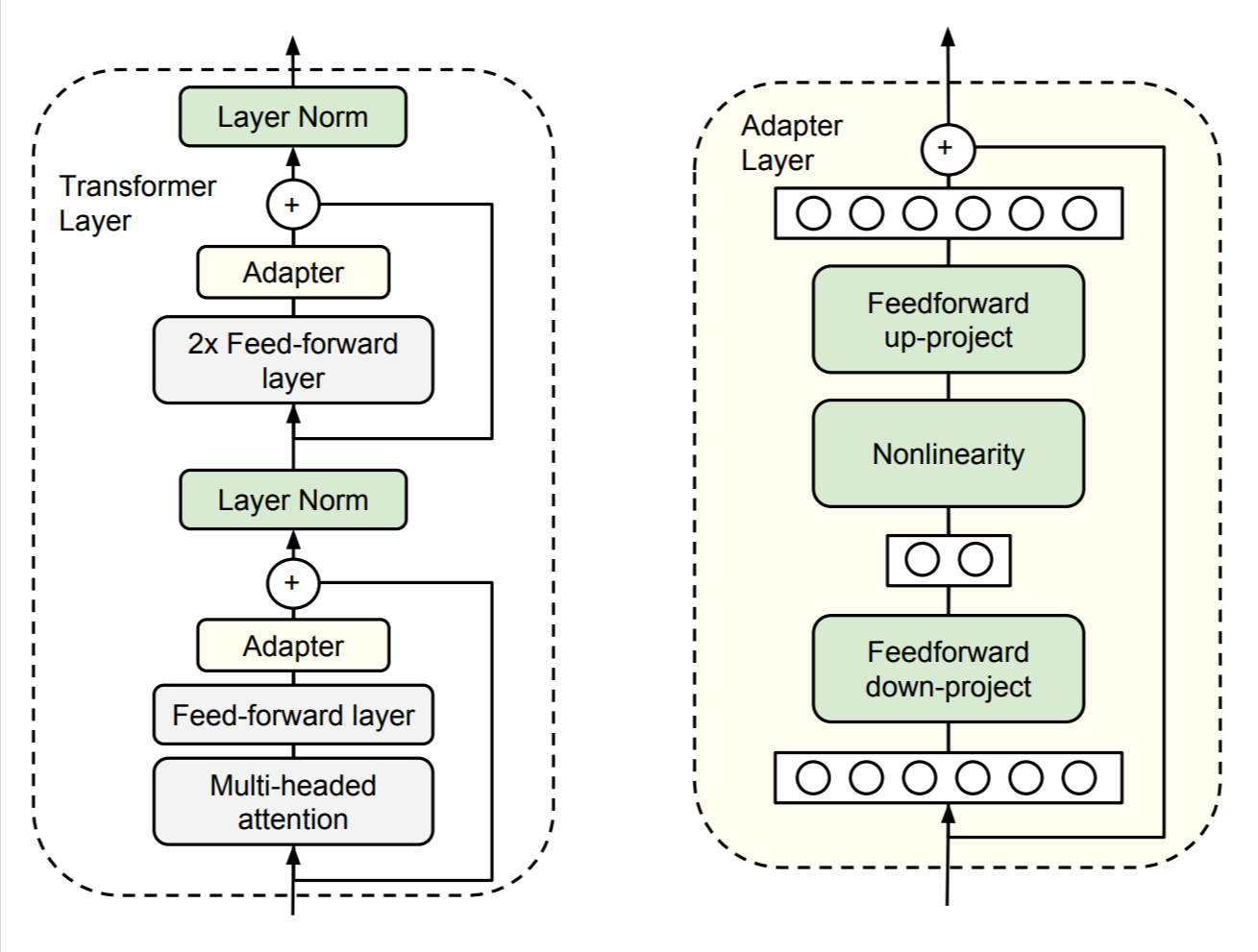
S. Houlsby, A. Giurgiu, S. Jastrzebski, B. Morrone, Q. de Laroussilhe, A. Gesmundo, M. Attariyan, S. Gelly
International Conference on Machine Learning 2019
paper
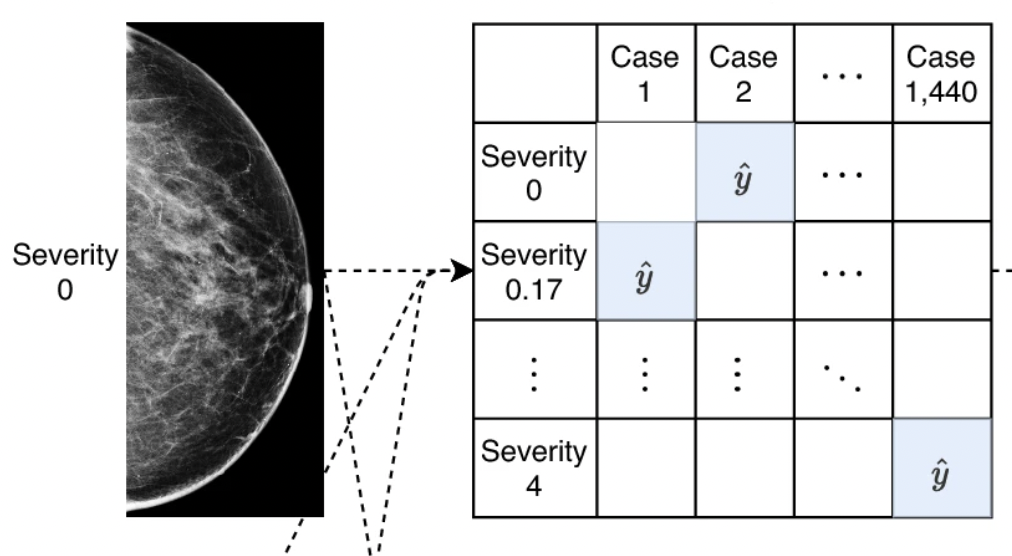
T. Makino, S. Jastrzebski, Witold Oleszkiewicz, [...], Kyunghyun Cho, Krzysztof J Geras
Nature Scientific Reports 2022
paper
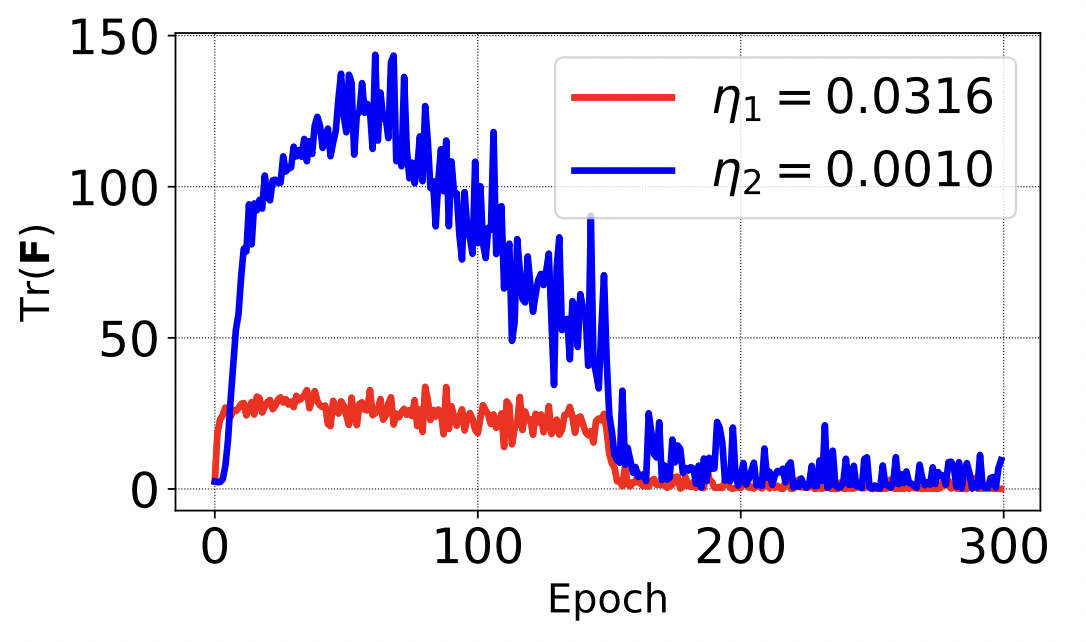
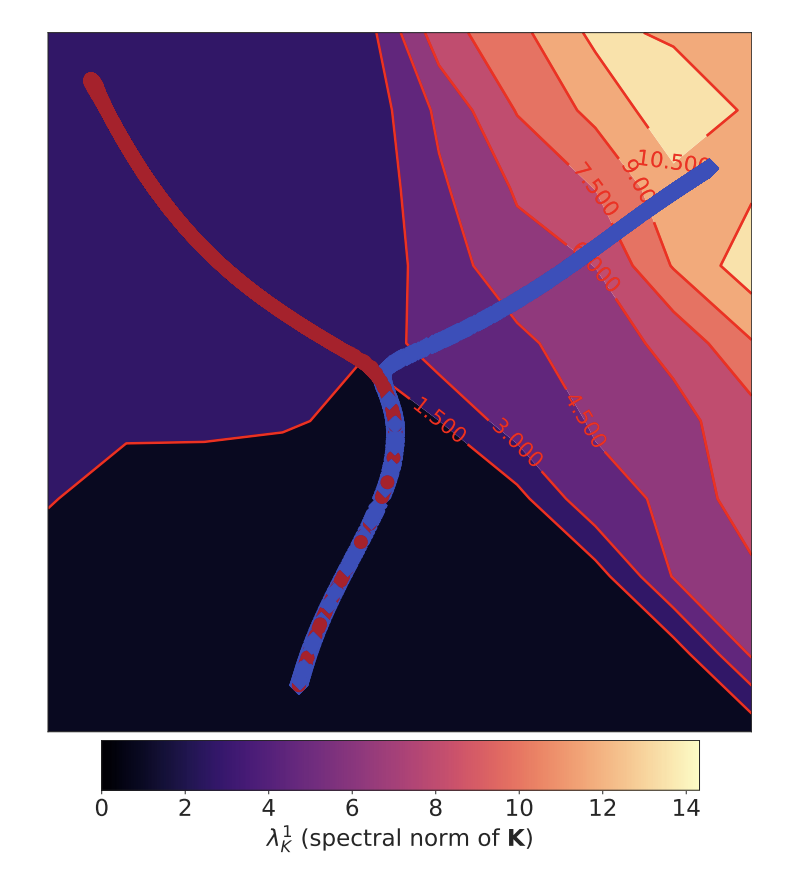
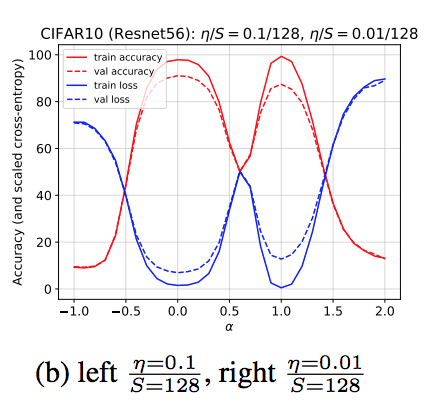
S. Jastrzębski*, Z. Kenton*, D. Arpit, N. Ballas, A. Fischer, Y. Bengio, A. Storkey
International Conference on Artificial Neural Networks 2018 (oral), International Conference on Learning Representations 2018 (workshop)
paper